genetools: single-cell analysis recipes (work in progress)¶
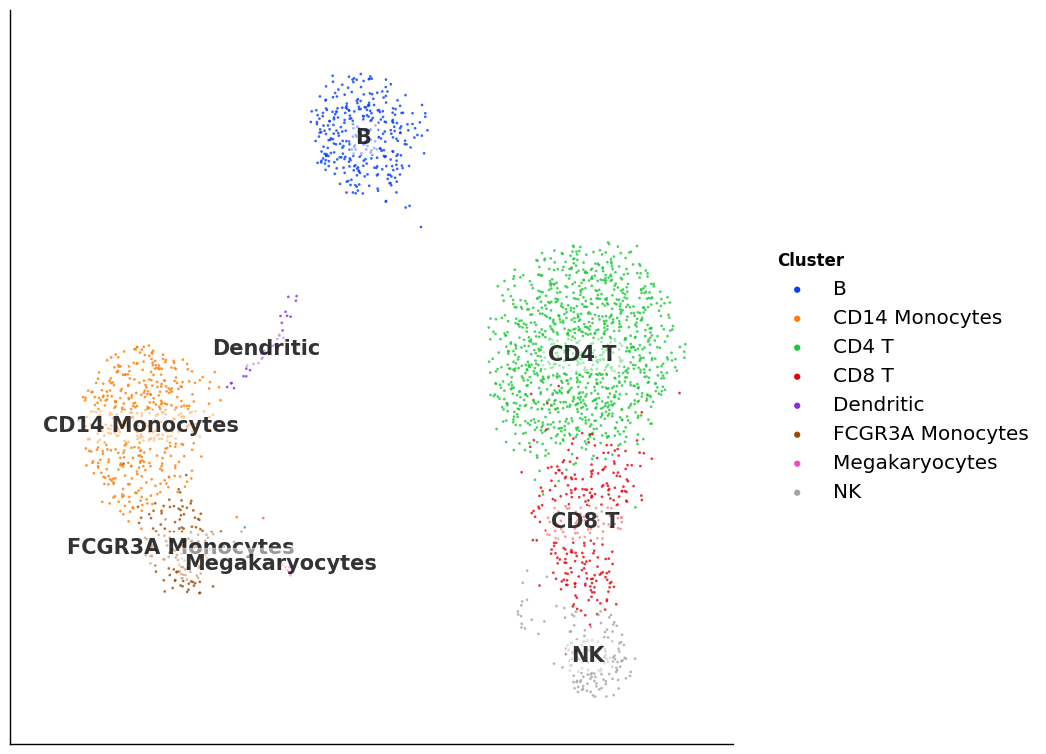
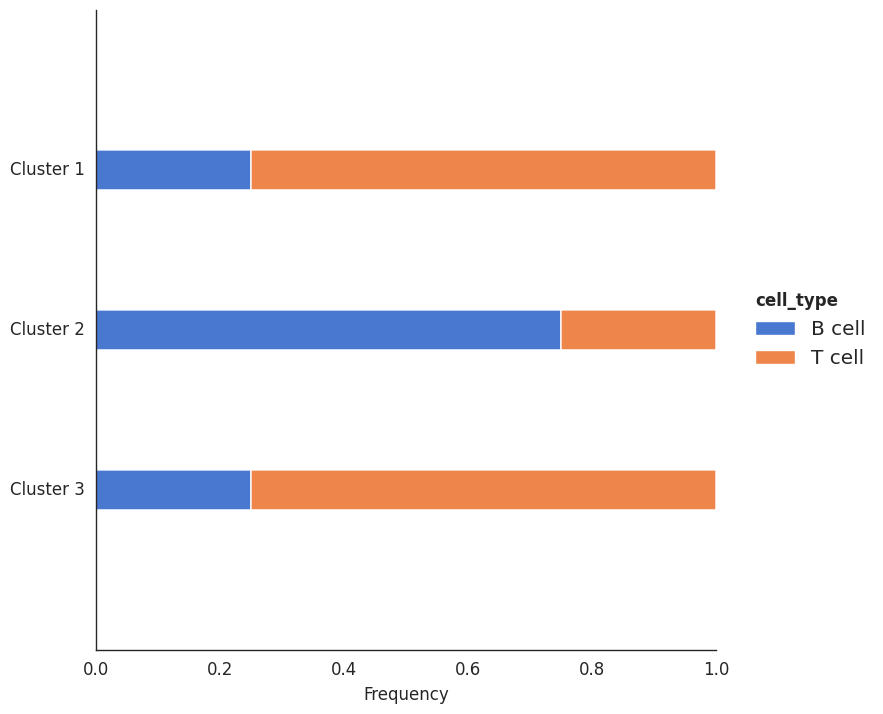
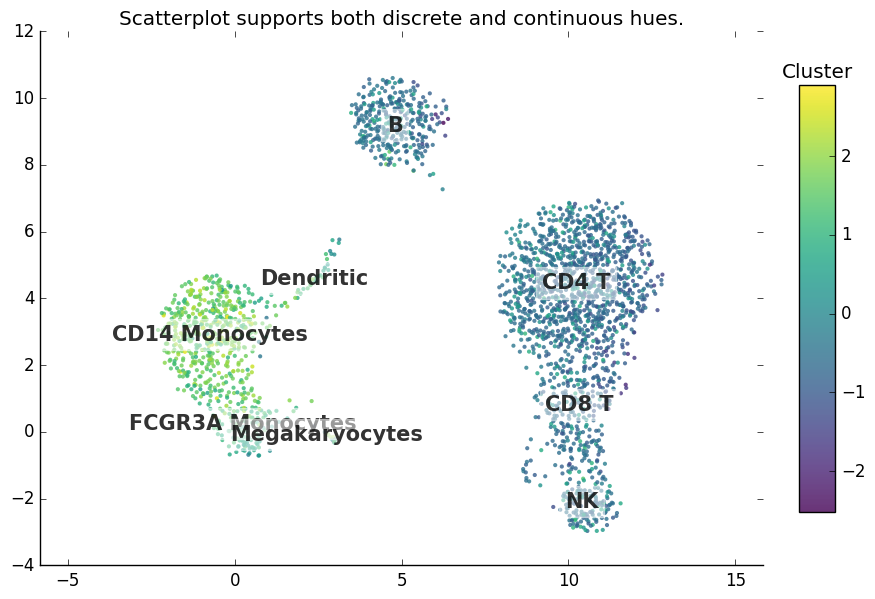
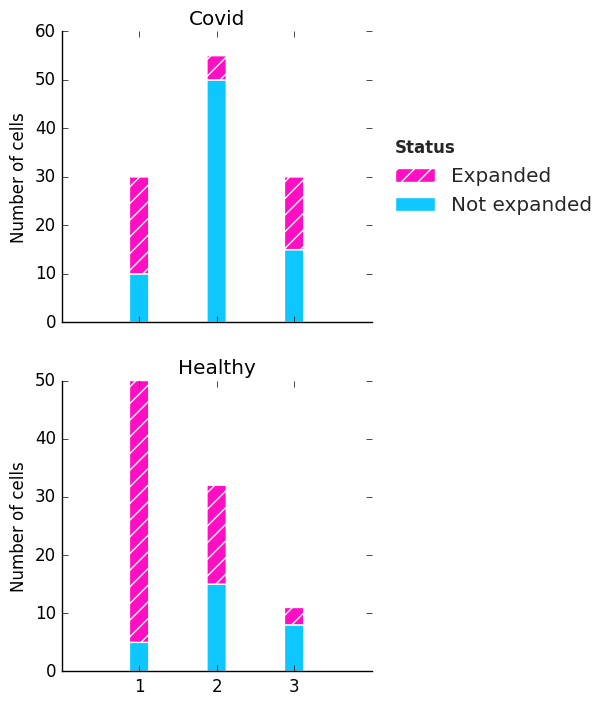
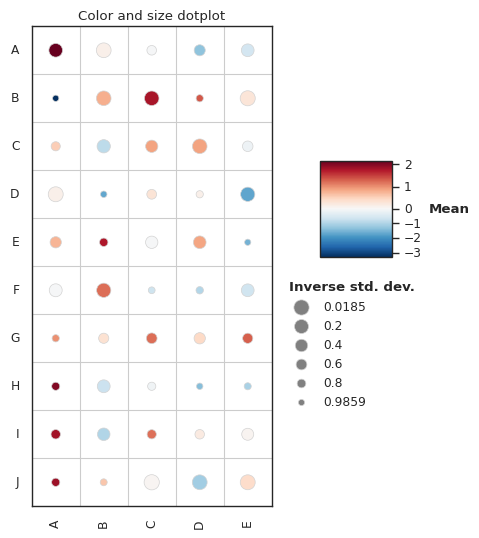
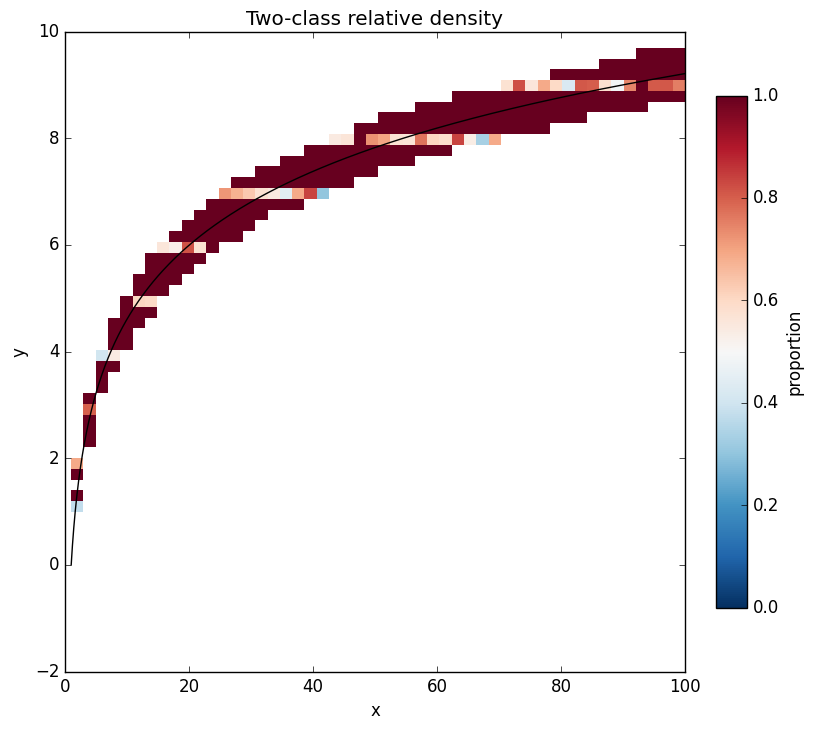
Plot gallery¶
Other features¶
Compare clustering results by computing co-clustering percentage.
Map marker genes against reference lists to find names for your clusters.
pandas shotrcuts:
Split single cell barcodes conveniently.
Defensive pandas merging and concatenation methods with strict correctness checks.
Full documentation: https://genetools.maximz.com.
Install¶
Run pip install --upgrade 'genetools[scanpy]'.
Or if you don’t use scanpy: pip install --upgrade genetools.
Usage¶
To use genetools in a project, add import genetools. Review the documentation and the tests for examples.
Development¶
Setup:
git clone git://github.com/maximz/genetools
cd genetools
pip install --upgrade pip wheel
pip install -r requirements_dev.txt
pre-commit install
Common commands:
# lint
make lint
# one-time: generate test anndata, and commit so we have reproducible tests in CI
rm -r data
make regen-test-data
# run tests locally
# this is done in a debian-based docker image to ensure image style matches what Github Actions CI will produce
# failing image snapshot tests are recorded in tests/results/
make build-docker-test-image # whenever requirements_dev.txt change
make test
# generate baseline figures (also happens in docker)
make regen-snapshot-figures
# regenerate test data, and baseline figures (also happens in docker)
make regen-test-data
# run tests locally without docker, therefore omitting the snapshot tests
# (the @snapshot_image tests are still executed but the images are not compared. the @pytest.mark.snapshot_custom are skipped altogether.)
make test-without-figures
# docs
make docs
# bump version before submitting a PR against master (all master commits are deployed)
bump2version patch # possible: major / minor / patch
# also ensure CHANGELOG.md updated
CI:
Main: Github Actions